Knn sklearn, K-Nearest Neighbor implementation with scikit learn

K-nearest neighbor implementation with scikit learn
Knn classifier implementation in scikit learn
In the introduction to k nearest neighbor and knn classifier implementation in Python from scratch, We discussed the key aspects of knn algorithms and implementing knn algorithms in an easy way for few observations dataset.
However in K-nearest neighbor classifier implementation in scikit learn post, we are going to examine the Breast Cancer Dataset using python sklearn library to model K-nearest neighbor algorithm. After modeling the knn classifier, we are going to use the trained knn model to predict whether the patient is suffering from the benign tumor or malignant tumor. The greatness of using Sklearn is that it provides us the functionality to implement machine learning algorithms in a few lines of code.
As we discussed the principle behind KNN classifier (K-Nearest Neighbor) algorithm is to find K predefined number of training samples closest in the distance to new point & predict the label from these. The distance measure is commonly considered to be Euclidean distance.
Euclidean distance
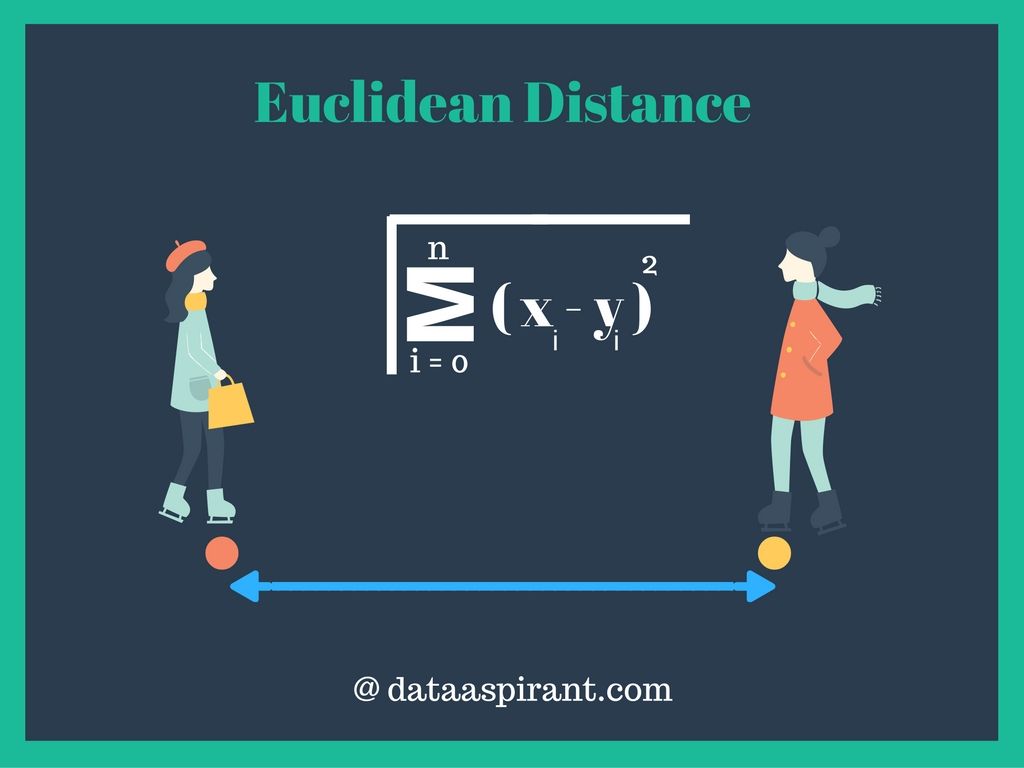




Euclidean Distance
Euclidean distance is the most commonly used distance measure. Euclidean distance also called as simply distance. The usage of Euclidean distance measure is highly recommended when data is dense or continuous. Euclidean distance is the best proximity measure. The Euclidean distance between two points is the length of the path connecting them.The Pythagorean theorem gives this distance between two points.
Euclidean distance implementation in python
#!/usr/bin/env python
from math import*
def euclidean_distance(x,y):
return sqrt(sum(pow(a-b,2) for a, b in zip(x, y)))
print euclidean_distance([0,3,4,5],[7,6,3,-1])
Script Output
9.74679434481 [Finished in 0.0s]
You can learn how to implement different similarity measure from the below post
KNN classifier is also considered to be an instance based learning / non-generalizing algorithm. It stores records of training data in a multidimensional space. For each new sample & particular value of K, it recalculates Euclidean distances and predicts the target class. So, it does not create a generalized internal model.
Similar to KNN classifier, we can use Radius Neighbor Classifier for classification tasks. As in KNN classifier, we specify the value of K, similarly, in Radius neighbor classifier the value of R should be defined. The RNC classifier determines the target class based on the number of neighbors within a fixed radius 



Knn implementation with Sklearn
Wisconsin Breast Cancer Data Set
The Wisconsin Breast Cancer Database was collected by Dr. William H. Wolberg (physician), University of Wisconsin Hospitals, USA. This dataset consists of 10 continuous attributes and 1 target class attributes. Class attribute shows the observation result, whether the patient is suffering from the benign tumor or malignant tumor. Benign tumors do not spread to other parts while the malignant tumor is cancerous. The dataset was collected & openly distributed so as to find out some patterns from this data.
Class attribute shows the observation result, whether the patient is suffering from the benign tumor or malignant tumor. Benign tumors do not spread to other parts while the malignant tumor is cancerous. The dataset was collected & openly distributed so as to find out some patterns from this data.


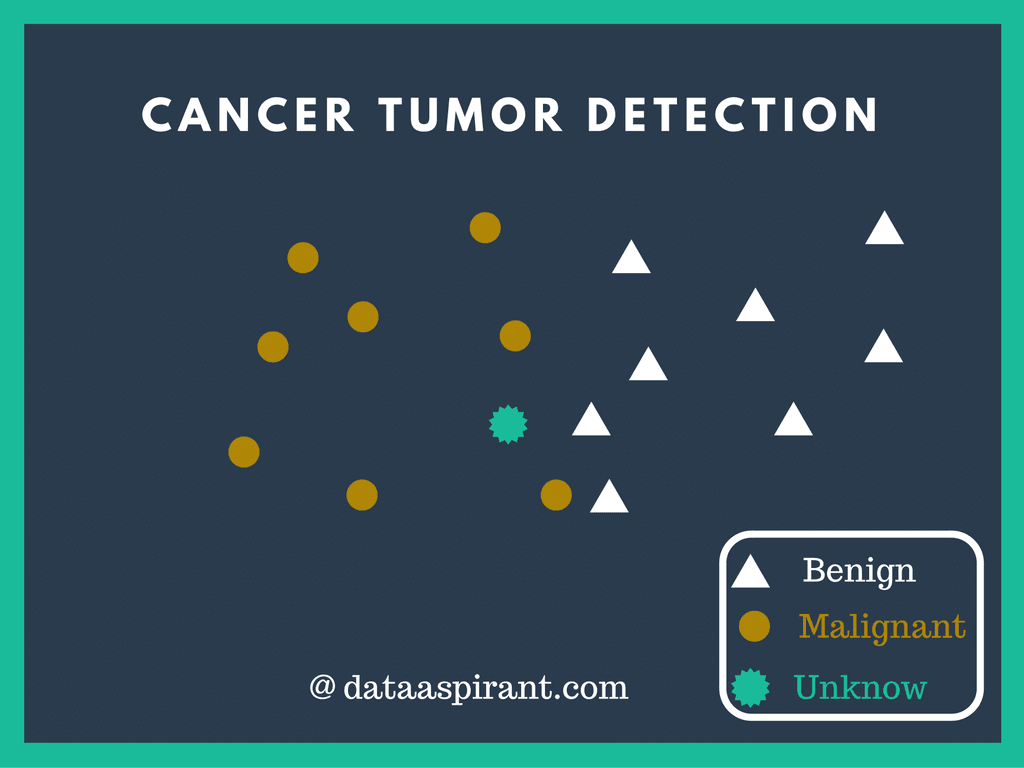

Cancer tumor detection with k-nearest neighbor
Breast Cancer Data Set Attribute Information:
1. Sample code number: id number
2. Clump Thickness: 1 – 10
3. Uniformity of Cell Size: 1 – 10
4. Uniformity of Cell Shape: 1 – 10
5. Marginal Adhesion: 1 – 10
6. Single Epithelial Cell Size: 1 – 10
7. Bare Nuclei: 1 – 10
8. Bland Chromatin: 1 – 10
9. Normal Nucleoli: 1 – 10
10. Mitoses: 1 – 10
11. Class: (2 for benign, 4 for malignant)
Problem Statement:
To model the knn classifier using the Breast Cancer data for predicting whether a patient is suffering from the benign tumor or malignant tumor.
KNN Model for Cancerous tumor detection:
To diagnose Breast Cancer, the doctor uses his experience by analyzing details provided by
- Patient’s Past Medical History
- Reports of all the tests performed.
Using the modeled KNN classifier, we will solve the problem in a way similar to the procedure used by doctors. The modeled KNN classifier will compare the new patient’s test reports, observation metrics with the records of patients(training data) that correctly classified as benign or malignant.
Python packages used:
- NumPy
- NumPy is a Numeric Python module. It provides fast mathematical functions.
- Numpy provides robust data structures for efficient computation of multi-dimensional arrays & matrices.
- We used numpy to read data files into numpy arrays and data manipulation.
- Scikit-Learn
- It’s a machine learning library. It includes various machine learning algorithms.
- We are using its
- Imputer,
- train_test_split,
- KNeighborsClassifier,
- accuracy_score algorithms.
If you haven’t setup the machine learning setup in your system the below posts will helpful.
K-nearest neighbor classifier implementation with scikit-learn
All the code snippets can be typed directly to jupyter Ipython notebook.
Libraries:
This section involves importing all the libraries. We are importing numpy and sklearn imputer, train_test_split, KneighborsClassifier & accuracy_score modules.
import numpy as np from sklearn.preprocessing import Imputer from sklearn.cross_validation import train_test_split from sklearn.neighbors import KNeighborsClassifier from sklearn.metrics import accuracy_score
Data Import:
We are using breast cancer data. You can download it from archive.ics.uci.edu website. For importing the data and manipulating it, we are going to use numpy arrays.
Using genfromtxt() method, we are importing our dataset into the 2d numpy array. You can import text files using this function. We are passing 3 parameters:
- fname
- It handles the filename with extension.
- delimiter
- The string used to separate values. In our dataset “,”(comma) is the separator.
- dtype
- It handles data type of variables.
All the values are numeric in our database. But some values are missing and are replaced by “?”. So, we will have to perform data imputation. Due to this reason, we are using float dtype.
cancer_data = np.genfromtxt( fname ='breast-cancer-wisconsin.data', delimiter= ',', dtype= float)
Using the above code we have imported our data into a 2d numpy array.
- len(): Function to find out the no. of records in our data.
- str(): Function to get an idea about the basic structure of data.
- shape: To get array dimensions.
print "Dataset Lenght:: ", len(cancer_data) print "Dataset:: ", str(cancer_data) print "Dataset Shape:: ", cancer_data.shape
Output:
Dataset Length:: 699
Dataset::
[[ 1.00002500e+06 5.00000000e+00 1.00000000e+00 ..., 1.00000000e+00
1.00000000e+00 2.00000000e+00]
[ 1.00294500e+06 5.00000000e+00 4.00000000e+00 ..., 2.00000000e+00
1.00000000e+00 2.00000000e+00]
[ 1.01542500e+06 3.00000000e+00 1.00000000e+00 ..., 1.00000000e+00
1.00000000e+00 2.00000000e+00]
...,
[ 8.88820000e+05 5.00000000e+00 1.00000000e+01 ..., 1.00000000e+01
2.00000000e+00 4.00000000e+00]
[ 8.97471000e+05 4.00000000e+00 8.00000000e+00 ..., 6.00000000e+00
1.00000000e+00 4.00000000e+00]
[ 8.97471000e+05 4.00000000e+00 8.00000000e+00 ..., 4.00000000e+00
1.00000000e+00 4.00000000e+00]]
Dataset Shape:: (699L, 11L)
The cancer dataset’s first column consists of patient’s id. To make this prediction process unbiased, we should remove this patient id. We can use numpy delete() method for this operation.
delete(): It returns a new transformed array. Three parameters should to passed.
- arr: It holds the array name.
- obj: It indicates which sub-arrays to remove.
- axis: The axis along which to delete. axis = 1 is used for columns & axis = 0 for rows.
cancer_data = np.delete(arr = cancer_data, obj= 0, axis = 1)
Now, we wish to divide the dataset into feature & label dataset. i.e., feature data is predictor variables they will help us to predict labels(criterion variable). Here, first 9 columns include continuous variables that will help us to predict whether a patient is having the benign tumor or malignant tumor.
X = cancer_data[:,range(0,9)] Y = cancer_data[:,9]
Data Imputation:
Imputation is a process of replacing missing values with substituted values. In our dataset, some columns have missing values. We can replace missing values with mean, median, mode or any particular value.
Sklearn provides Imputer() method to perform imputation in 1 line of code. We just need to define missing_values, axis, and strategy. We are using “median” value of the column to substitute with the missing value.
imp = Imputer(missing_values="NaN", strategy='median', axis=0) X = imp.fit_transform(X)
Train, Test data split:
For dividing data into train data & test data. We are using train_test_split() method by sklearn.
train_test_split(): We are using 4 parameters X, Y, test_size, random_state
- X, Y: X is a numpy array consisting of feature dataset & Y contains labels for each record.
- test_size: It represents the size of test data needs to split. If we use 0.4, it indicates 40% of data should be separated and saved as testing data.
- random_state: It’s pseudo-random number generator state used for random sampling. If you want to replicate our results, then use the same value of random_state.
Now, X_train & y_train are training datasets. X_test & y_test are testing datasets.
y_train & y_test are 2d numpy arrays with 1 column. To convert it into a 1d array, we are using ravel().
X_train, X_test, y_train, y_test = train_test_split( X, Y, test_size = 0.3, random_state = 100) y_train = y_train.ravel() y_test = y_test.ravel()
KNN Implementation:
Now we are fitting KNN algorithm on training data, predicting labels for dataset and printing the accuracy of the model for different values of K(ranging from 1 to 25).
KNeighborsClassifier(): This is the classifier function for KNN. It is the main function for implementing the algorithms. Some important parameters are:
- n_neighbors: It holds the value of K, we need to pass and it must be an integer. If we don’t give the value of n_neighbors then by default, it takes the value as 5.
- Weights: It holds a string value i.e., name of the weight function. The Weight function used in prediction. It can hold values like ‘uniform’ or ‘distance’ or any user defined function.
- ‘uniform’ weight used when all points in the neighborhood are weighted equally. Default value for weights taken as ‘uniform’
- ‘distance’ weight used for giving closer neighbors- higher weight and far neighbors-less weight, i.e., weight points by the inverse of their distance.
- user defined function we can call the user defined functions. The user defined function can used when we want to produce custom weight values. It accepts distance values and returns an array of weights.
- algorithm: It specifies algorithm which should be used to compute the nearest neighbors. It can values like ‘auto’, ‘ball_tree’, ‘kd_tree’, brute’. It is an optional parameter.
- a) ‘ball_tree’ , ‘kd_tree’ are used to implement ball tree algorithm. These are special kind of data structures for space partitioning.
- b) ‘brute’ is used to implement brute-force search algorithm.
- c) ‘auto’ is used to give control to the system. By using ‘auto’, it automatically decides the best algorithm according to values of training data.fit()
- data.fit(): A fit method is used to fit the model. It is passed with two parameters:X and Y. For training data fitting on KNN algorithm, this needs to call.
- X: It consists of training data with features.
- Y: It consists of training data with labels.predict(): It predicts class labels for the data provided as its parameters.
If an array of features data is entered as parameters, then an array of labels is given as output.
Accuracy Score:
accuracy_score(): This function is used to print accuracy of KNN algorithm. By accuracy, we mean the ratio of the correctly predicted data points to all the predicted data points. Accuracy as a metric helps to understand the effectiveness of our algorithm. It takes 4 parameters.
- y_true,
- y_pred,
- normalize,
- sample_weight.
Out of these 4, normalize & sample_weight are optional parameters. The parameter y_true accepts an array of correct labels and y_pred takes an array of predicted labels that are returned by the classifier. It returns accuracy as a float value.
for K in range(25): K_value = K+1 neigh = KNeighborsClassifier(n_neighbors = K_value, weights='uniform', algorithm='auto') neigh.fit(X_train, y_train) y_pred = neigh.predict(X_test) print "Accuracy is ", accuracy_score(y_test,y_pred)*100,"% for K-Value:",K_value
Output:
Accuracy is 95.2380952381 % for K-Value: 1 Accuracy is 93.3333333333 % for K-Value: 2 Accuracy is 95.7142857143 % for K-Value: 3 Accuracy is 95.2380952381 % for K-Value: 4 Accuracy is 95.7142857143 % for K-Value: 5 Accuracy is 94.7619047619 % for K-Value: 6 Accuracy is 94.7619047619 % for K-Value: 7 Accuracy is 94.2857142857 % for K-Value: 8 Accuracy is 94.7619047619 % for K-Value: 9 Accuracy is 94.2857142857 % for K-Value: 10 Accuracy is 94.2857142857 % for K-Value: 11 Accuracy is 94.7619047619 % for K-Value: 12 Accuracy is 94.7619047619 % for K-Value: 13 Accuracy is 93.8095238095 % for K-Value: 14 Accuracy is 93.8095238095 % for K-Value: 15 Accuracy is 93.8095238095 % for K-Value: 16 Accuracy is 93.8095238095 % for K-Value: 17 Accuracy is 93.8095238095 % for K-Value: 18 Accuracy is 93.8095238095 % for K-Value: 19 Accuracy is 93.8095238095 % for K-Value: 20 Accuracy is 93.8095238095 % for K-Value: 21 Accuracy is 93.8095238095 % for K-Value: 22 Accuracy is 93.8095238095 % for K-Value: 23 Accuracy is 93.8095238095 % for K-Value: 24 Accuracy is 93.8095238095 % for K-Value: 25
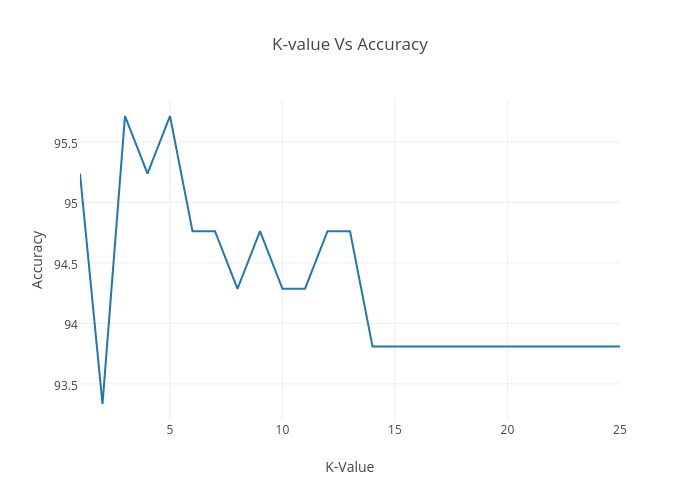
K value Vs Accuracy Change Graph
It shows that we are getting 95.71% accuracy on K = 3, 5. Choosing a large value of K will lead to greater amount of execution time & underfitting. Selecting the small value of K will lead to overfitting. There is no such guaranteed way to find the best value of K. So, to run it quickly we are considering K =3 for this tutorial.
Further reading
- Manipulating parameters of KNeighborsClassifier() method.
- Implementing KNN on cross-validation data. It helps to validate which K value may give the better accuracy.
Follow us:
FACEBOOK| QUORA |TWITTER| GOOGLE+ | LINKEDIN| REDDIT | FLIPBOARD | MEDIUM | GITHUB
I hope you like this post. If you have any questions, then feel free to comment below. If you want me to write on one particular topic, then do tell it to me in the comments below.
Related Courses:
Do check out unlimited data science courses
| Title of the course | Course Link | Course Link |
Machine Learning: Classification    |
Machine Learning: Classification |
|
Data Mining with Python: Classification and Regression    |
Data Mining with Python: Classification and Regression |
|
Machine learning with Scikit-learn    |
Machine learning with Scikit-learn |
|





Thanks, great article. I was trying to run the same on my machine and faced with this issue. Do you have any pointers?
X = imp.fit_transform(X)
File “/usr/lib/python2.7/dist-packages/sklearn/base.py”, line 408, in fit_transform
return self.fit(X, **fit_params).transform(X)
File “/usr/lib/python2.7/dist-packages/sklearn/preprocessing/imputation.py”, line 193, in fit
self.axis)
File “/usr/lib/python2.7/dist-packages/sklearn/preprocessing/imputation.py”, line 307, in _dense_fit
median[np.ma.getmask(median_masked)] = np.nan
IndexError: in the future, 0-d boolean arrays will be interpreted as a valid boolean index
Hi Vikas,
Thanks for your compliment. Could please check the installing Python machine learning libraries before running the code. If you face any issues let us know.