Five most popular similarity measures implementation in python
Five most popular similarity measures implementation in python
The buzz term similarity distance measure or similarity measures has got a wide variety of definitions among the math and machine learning practitioners. As a result, those terms, concepts, and their usage went way beyond the minds of the data science beginner. Who started to understand them for the very first time.
So today we wrote this post to give more clear and very intuitive definitions for similarity. We will also drive you to the five most popular similarity measures and the implementation of them in the python programming language.
Before going to explain different similarity distance measures. Let me explain the effective key term similarity in data mining or machine learning.
This similarity is the very basic building block for activities such as
- Recommendation engines,
- clustering,
- Different classification problems,
- Email spam or ham classification problems
Before we drive further, below are the topics you will be learning in this article.
Table of contents
- Similarity
- Euclidean distance
- Euclidean distance implementation in python
- Manhattan distance
- Manhattan distance implementation in python
- Minkowski distance
- Synonyms of Minkowski
- Minkowski distance implementation in python
- Cosine Similarity
- Cosine Similarity Implementation In Python
- Jaccard Similarity
- Sets & Set Operations
- Jaccard Similarity implementation in python
- Implementations of all five similarity measures implementation in python
Similarity
The similarity measure is the measure of how much alike two data objects are. A similarity measure is a data mining or machine learning context is a distance with dimensions representing features of the objects. If the distance is small, the features are having a high degree of similarity. Whereas a large distance will be a low degree of similarity.
Similarity measure usage is more in the text related preprocessing techniques, Also the similarity concepts used in advanced word embedding techniques. We can use these concepts in various deep learning applications. Uses the difference between the image for checking the data created with data augmentation techniques.
The similarity is subjective and is highly dependent on the domain and application.
For example, two fruits are similar because of color or size or taste. Special care should be taken when calculating distance across dimensions/features that are unrelated. The relative values of each element must be normalized, or one feature could end up dominating the distance calculation.
Generally, similarity are measured in the range 0 to 1 [0,1]. In the machine learning world, this score in the range of [0, 1] is called the similarity score.
Two main consideration of similarity:
- Similarity = 1 if X = Y (Where X, Y are two objects)
- Similarity = 0 if X ≠ Y
That’s all about similarity let’s drive to five most popular similarity distance measures.
Euclidean distance
Euclidean distance is the most common use of distance measure. In most cases when people say about distance, they will refer to Euclidean distance.
Euclidean distance is also known as simply distance. When data is dense or continuous, this is the best proximity measure.
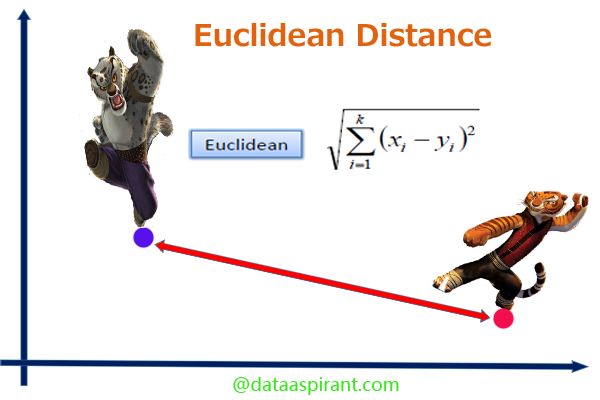
The Euclidean distance between two points is the length of the path connecting them. The Pythagorean theorem gives this distance between two points.
Euclidean distance implementation in python:
#!/usr/bin/env python
from math import*
def euclidean_distance(x,y):
return sqrt(sum(pow(a-b,2) for a, b in zip(x, y)))
print euclidean_distance([0,3,4,5],[7,6,3,-1])
Script Output:
9.74679434481 [Finished in 0.0s]
Manhattan distance:
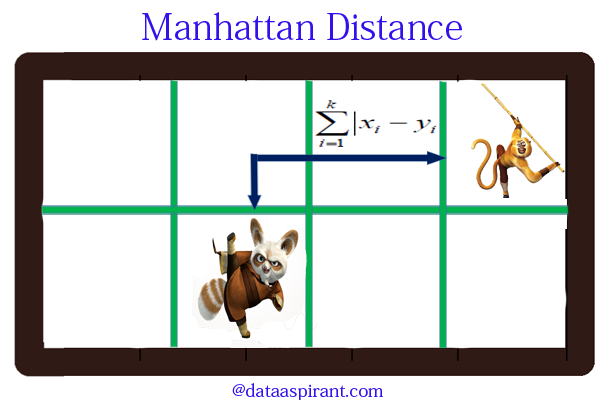
Manhattan distance is a metric in which the distance between two points is calculated as the sum of the absolute differences of their Cartesian coordinates. In a simple way of saying it is the total sum of the difference between the x-coordinates and y-coordinates.
Suppose we have two points A and B. If we want to find the Manhattan distance between them, just we have, to sum up, the absolute x-axis and y-axis variation. This means we have to find how these two points A and B are varying in X-axis and Y-axis. In a more mathematical way of saying Manhattan distance between two points measured along axes at right angles.
In a plane with p1 at (x1, y1) and p2 at (x2, y2).
Manhattan distance = |x1 – x2| + |y1 – y2|
This Manhattan distance metric is also known as Manhattan length, rectilinear distance, L1 distance or L1 norm, city block distance, Minkowski’s L1 distance, taxi-cab metric, or city block distance.
Manhattan distance implementation in python:
#!/usr/bin/env python
from math import*
def manhattan_distance(x,y):
return sum(abs(a-b) for a,b in zip(x,y))
print manhattan_distance([10,20,10],[10,20,20])
Script Output:
10 [Finished in 0.0s]
Minkowski distance
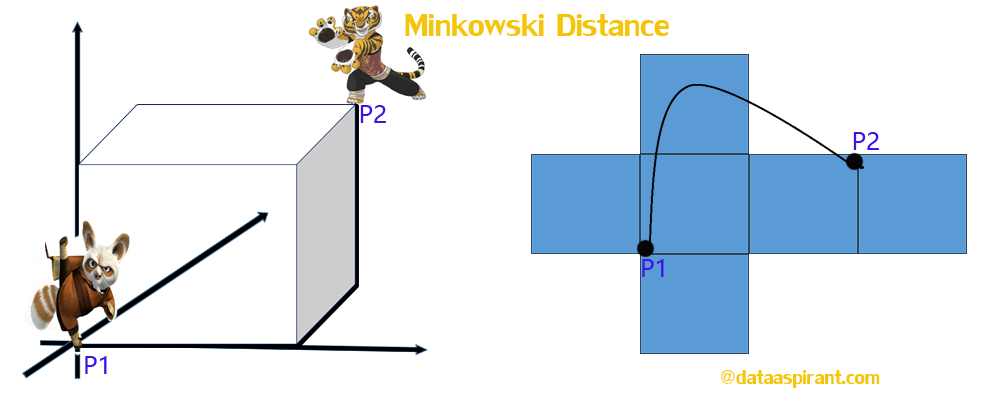
The Minkowski distance is a generalized metric form of Euclidean distance and Manhattan distance.
In the equation, d^MKD is the Minkowski distance between the data record i and j, k the index of a variable, n the total number of variables y and λ the order of the Minkowski metric. Although it is defined for any λ > 0, it is rarely used for values other than 1, 2, and ∞.
The way distances are measured by the Minkowski metric of different orders between two objects with three variables ( In the image is displayed in a coordinate system with x, y, z-axes).
Synonyms of Minkowski
Different names for the Minkowski distance or Minkowski metric arise from the order:
- λ = 1 is the Manhattan distance. Synonyms are L1-Norm, Taxicab, or City-Block distance. For two vectors of ranked ordinal variables, the Manhattan distance is sometimes called Foot-ruler distance.
- λ = 2 is the Euclidean distance. Synonyms are L2-Norm or Ruler distance. For two vectors of ranked ordinal variables, the Euclidean distance is sometimes called Spear-man distance.
- λ = ∞ is the Chebyshev distance. Synonyms are Lmax-Norm or Chessboard distance.
reference.
Minkowski distance implementation in python
#!/usr/bin/env python
from math import*
from decimal import Decimal
def nth_root(value, n_root):
root_value = 1/float(n_root)
return round (Decimal(value) ** Decimal(root_value),3)
def minkowski_distance(x,y,p_value):
return nth_root(sum(pow(abs(a-b),p_value) for a,b in zip(x, y)),p_value)
print minkowski_distance([0,3,4,5],[7,6,3,-1],3)
Script Output:
8.373 [Finished in 0.0s]
Cosine Similarity
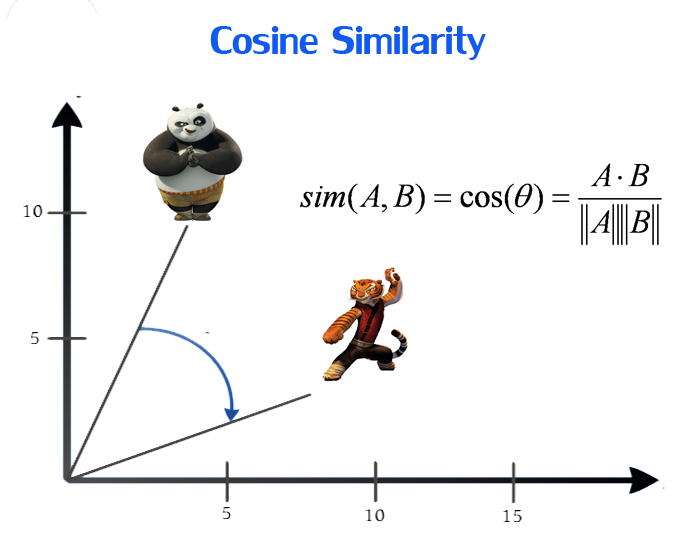
The cosine similarity metric finds the normalized dot product of the two attributes. By determining the cosine similarity, we would effectively try to find the cosine of the angle between the two objects. The cosine of 0° is 1, and it is less than 1 for any other angle.
It is thus a judgment of orientation and not magnitude. Two vectors with the same orientation have a cosine similarity of 1, two vectors at 90° have a similarity of 0. Whereas two vectors diametrically opposed having a similarity of -1, independent of their magnitude.
Cosine similarity is particularly used in positive space, where the outcome is neatly bounded in [0,1]. One of the reasons for the popularity of cosine similarity is that it is very efficient to evaluate, especially for sparse vectors.
Cosine similarity implementation in python
#!/usr/bin/env python
from math import*
def square_rooted(x):
return round(sqrt(sum([a*a for a in x])),3)
def cosine_similarity(x,y):
numerator = sum(a*b for a,b in zip(x,y))
denominator = square_rooted(x)*square_rooted(y)
return round(numerator/float(denominator),3)
print cosine_similarity([3, 45, 7, 2], [2, 54, 13, 15])
Script Output:
0.972 [Finished in 0.1s]
Jaccard similarity:
So far discussed some metrics to find the similarity between objects. where the objects are points or vectors. When we consider Jaccard similarity these objects will be sets.
So first let’s learn some very basics about sets and set operations
Sets & Set Operations
Sets
A set is (unordered) collection of objects {a,b,c}. we use the notation as elements separated by commas inside curly brackets { }. They are unordered so {a,b} = { b,a }.
Cardinality
The cardinality of A denoted by |A| which counts how many elements are in A.
Intersection
The intersection between two sets A and B is denoted A ∩ B and reveals all items which are in both sets A, B.
Union
The union between two sets A and B is denoted A ∪ B and reveals all items which are in either set.
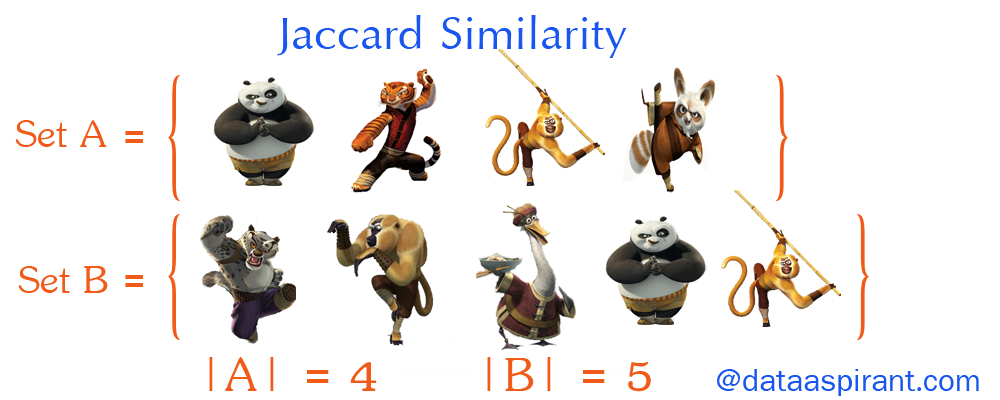
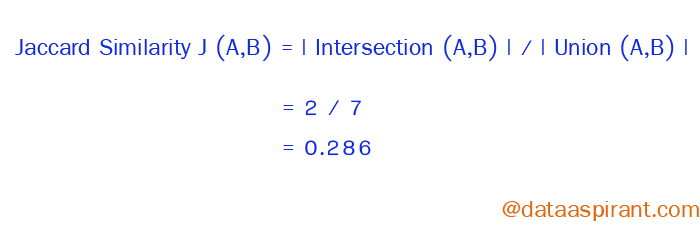
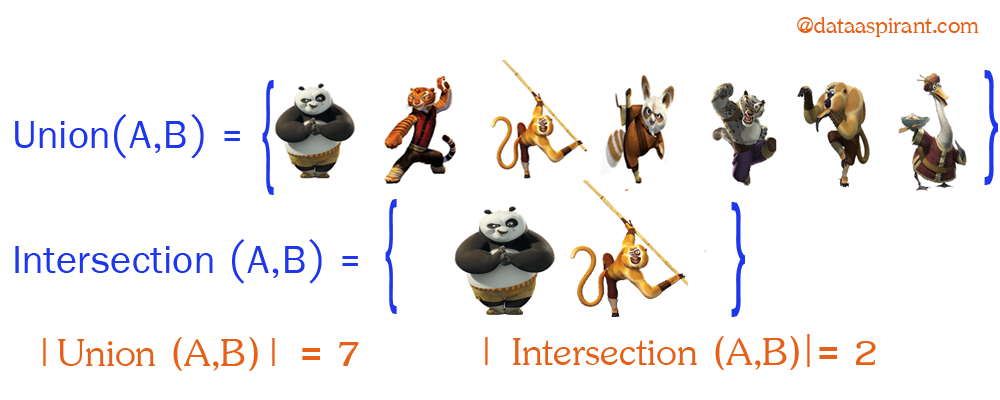
Now going back to Jaccard similarity. The Jaccard similarity measures the similarity between finite sample sets and is defined as the cardinality of the intersection of sets divided by the cardinality of the union of the sample sets.
Suppose you want to find Jaccard similarity between two sets A and B it is the ration of the cardinality of A ∩ B and A ∪ B
Jaccard similarity implementation in Python
#!/usr/bin/env python
from math import*
def jaccard_similarity(x,y):
intersection_cardinality = len(set.intersection(*[set(x), set(y)]))
union_cardinality = len(set.union(*[set(x), set(y)]))
return intersection_cardinality/float(union_cardinality)
print jaccard_similarity([0,1,2,5,6],[0,2,3,5,7,9])
Script Output:
0.375 [Finished in 0.0s]
Implementation of all five similarity measure into one Similarity class
file_name : similaritymeasures.py
#!/usr/bin/env python
from math import*
from decimal import Decimal
class Similarity():
""" Five similarity measures function """
def euclidean_distance(self,x,y):
""" return euclidean distance between two lists """
return sqrt(sum(pow(a-b,2) for a, b in zip(x, y)))
def manhattan_distance(self,x,y):
""" return manhattan distance between two lists """
return sum(abs(a-b) for a,b in zip(x,y))
def minkowski_distance(self,x,y,p_value):
""" return minkowski distance between two lists """
return self.nth_root(sum(pow(abs(a-b),p_value) for a,b in zip(x, y)),
p_value)
def nth_root(self,value, n_root):
""" returns the n_root of an value """
root_value = 1/float(n_root)
return round (Decimal(value) ** Decimal(root_value),3)
def cosine_similarity(self,x,y):
""" return cosine similarity between two lists """
numerator = sum(a*b for a,b in zip(x,y))
denominator = self.square_rooted(x)*self.square_rooted(y)
return round(numerator/float(denominator),3)
def square_rooted(self,x):
""" return 3 rounded square rooted value """
return round(sqrt(sum([a*a for a in x])),3)
def jaccard_similarity(self,x,y):
""" returns the jaccard similarity between two lists """
intersection_cardinality = len(set.intersection(*[set(x), set(y)]))
union_cardinality = len(set.union(*[set(x), set(y)]))
return intersection_cardinality/float(union_cardinality)
Using Similarity class
#!/usr/bin/env python
from similaritymeasures import Similarity
def main():
""" the main function to create Similarity class instance and get used to it """
measures = Similarity()
print measures.euclidean_distance([0,3,4,5],[7,6,3,-1])
print measures.jaccard_similarity([0,1,2,5,6],[0,2,3,5,7,9])
if __name__ == "__main__":
main()
You can get all the complete codes of dataaspirant at dataaspirant data science codes
Related Courses
Do check out unlimited data science courses
- Introduction to natural language processing
- Natural language processing specialization course
- Nlp Modules building with python
Follow us:
FACEBOOK| QUORA |TWITTER| GOOGLE+ | LINKEDIN| FLIPBOARD | MEDIUM | GITHUB
I hope you like this post. If you have any questions then feel free to comment below. If you want me to write on one specific topic then do tell it to me in the comments below.


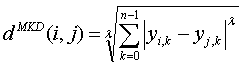







I have read many data science posts online previously, but none has managed to captivate my attention like this one. This is truly a masterpiece, and a perfect guide for all data science aspirants. Thanks to the writer for spelling out the concepts clearly, and using just the right words and structure.
Wow, your words truly warmed my heart! It’s immensely rewarding to hear such positive feedback. My aim is always to make complex concepts accessible and engaging for readers like you. Knowing that this post resonated and provided value makes all the effort worth it. Thank you for your encouragement, and if you ever have any questions or topics you’d like to delve into, please reach out. Here’s to more learning and discovery in data science! 🌟📊
I wonder if there is a way to find similarity measure both magnitude and direction? for example, let we have n vectors, but I want to select m vectors such that m<n, such that m can represent the n population. one way could be finding the projection of each vector against each other, this will give us direction, and then we look into the magnitude using any method explain in this article. then we take the intersection of both the projections and magnitude similarity.
I don't know if I explain the point correctly.
Hello it’s me, I am also visiting this site regularly, this website is truly pleasant and the people are really sharing good thoughts.
Thanks! 🙂
great publish, very informative. I ponder why the other specialists of this sector don’t
understand this. You must continue your writing.
I am sure, you have a huge readers’ base already!
Thanks! Sallie Grandi 🙂
I think this is one of the such a lot important info for me.
And i am glad studying your article. However wanna remark
on some basic issues, The website style is great, the articles is actually great : D.
Just right job, cheers
Thanks! for the compliment. 🙂
Hi there! This post couldn’t be written anyy better! Reading through this post reminds me of my good old room
mate! He always kept chatting about this. I will forward this write-up to him.
Pretty sure he wilkl have a good read. Thank you
for sharing!
Thanks, Demetra Barger,
We are glad about this, we wish you happy learning!
i am searching for similarity measure using correlation ?can anyone help me about this
Hi Vinod Kumar,
You can check the person’s similairy measure in this article please have a look.
Thanks,
Saimadhu
proximity=1-1/1+e^-(u1-u2)
In users-items matric ..how to write this formula in python code
Hi Fawad Ahmad Qureshi,
You can use NumPy to write code in a much similar way for this use case. Please check once, If you find it difficult, let me know, will share with you the code.
Thanks,
Saimadhu
very helpful!
Hi Dr kim,
Thanks for the compliment.
We wish you a very happy learning.
Good post
However multiple grammatical errors
Would love to correct them and contribute towards the site
Hi Chaitanya Bapat,
I am glad for your suggestion. IF you would like to contribute content please join us.
Excellent work bro. I love you.
Thanks 🙂
Nice Post.Thanks a lot.:)
Nice Post It is easily understood with list of x and y (two lists). so please I want to know more how to implement for large documents especially for cosine similarity in IR
Hi Misgu,
Thanks for your compliment.
fantastic images
Thank you 🙂
Your post seems to cover just one similarity measure: Jaccard. The remaining four are distance metrics; they must be transformed to provide similarity.
I actually found Jaccard’s metric to work nicely for weighted sets as well: if an item occurs in both A and B, its weight in the intersection is the minimum of the two weights, and its weight in the union is the maximum of the two weights.
Reblogged this on Random and commented:
A good blog, explaining some important similarity metrics.
I don’t think there is no need to write your own implementation. All of them and a lot more are already available in scipy.spatial.distance module in python.
Hi Jitesh Khandelwal!
You said true, but we have to explain how we can implement them.
Thought you might cover Mahalanobis distance.
Agreed … Mahalanobis distance and Haversine distance are missing .. I do not know of any application of Minowski distance ( for lambda >2) … (except Chebyshev )
Agreed … Mahalanobis distance and Haversine distance are missing … I dont know of any application of Minowski distance for lambda > 2 (except Chebyshev)
A few points:
1. A measure of similarity need not be symmetrical
2. A measure of similarity is not a metric space
3. Information theoretic measures, like KL and Mutual Information tend to be the most powerful, but the most difficult to manipulate mathematically.